 
Improved resolution of minor shifts in buoyant density relative to TRFLP analysis alone was confirmed using well-controlled SIP analyses. The use of multiple restriction enzymes in terminal restriction fragment length polymorphism analysis and identification of performance-related caecal. 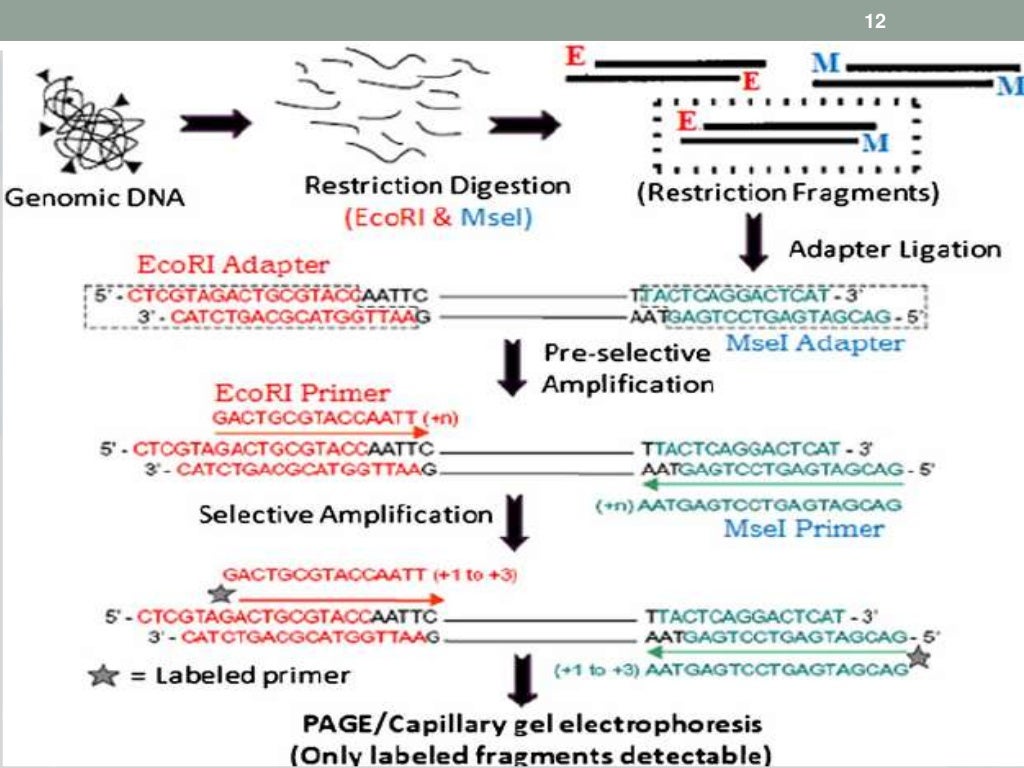
Method accuracy was evaluated using mixtures of cloned 16S rRNA genes, DNA extracted from low- and high-G+C bacterial isolates (Escherichia coli, Rhodococcus, Variovorax, and Microbacterium), and DNA from soil microcosms amended with known amounts of genomic DNA from bacterial isolates. This method combines well-controlled amplification with TRFLP analysis to quantify relative taxon abundance in amplicon pools of FAM-labeled PCR products, using the intercalating dye EvaGreen to monitor amplification. RFLP (often pronounced rif lip, as if it were a word) is a method used by molecular biologists to follow a particular sequence of DNA as it is passed on. Since then several modified protocols have been reported, but all typically include five main steps: (a) restriction of genomic DNA (see Note 1) and ligation of adaptors (most often performed together) to restricted fragments (b) preselective PCR amplification of a subset of the restricted fragments (c) selective PCR amplification, reducing fu. We therefore developed a tandem quantitative PCR (qPCR)-TRFLP (terminal restriction fragment length polymorphism) protocol that improves resolution of detection by quantifying specific taxonomic groups in gradient fractions. Restriction fragment length polymorphism analysis This unit describes Restriction Fragment Length Polymorphism (RFLP) analysis, which utilizes restriction endonuclease digestion to identify DNA sequence polymorphisms in genes or DNA regions of interest. However, application of SIP to microbial populations with relatively minor buoyant density increases, such as populations that utilize compounds as a nitrogen source, results in reduced resolution of labeled populations. Such differences were originally detected as variations in the length of restriction fragments (restriction fragment length polymorphisms or RFLPs) after. Restriction fragment length polymorphism (RFLP) (pronounced rif lip) is one of the earliest molecular markers developed for genetic mapping and one of the. detection of point mutations after the genomic sequences are amplified by PCR. 
Stable-isotope probing (SIP) has proved a valuable cultivation-independent tool for linking specific microbial populations to selected functions in various natural and engineered systems. reaction-restriction fragment length polymorphism (PCR-RFLP) method, for the detection of single nucleotide polymorphisms (SNPs). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
